From backyard projects to commercial bat surveys, the BTO Acoustic Pipeline provides tools for detecting and identifying birds, bats and other wildlife in audible and ultrasonic sound recordings. See our about passive acoustic monitoring and BTO’s acoustic monitoring research pages to learn more about acoustic monitoring. If you'd like to help us further unlock the potential of acoustic monitoring for birds and develop even more effective monitoring tools, please consider donating to our Acoustic Appeal.

About the Pipeline
About the The BTO Acoustic Pipeline enables accurate species identification and data management for acoustic monitoring, in conservation, management and site assessment. It offers bespoke audible classifiers for several key bird species and groups, and ultrasonic classifiers for bats, small mammals, bush-crickets and moths.
It is used across the globe in conservation, management and site assessment, providing a smooth and easy end-to-end workflow, with free support from our in-house experts. Free or scalable paid plans with bespoke data management options are available.
- Get in touch at acoustic.pipeline@bto.org or get started today.
How it works
1. Download the Acoustic Pipeline App to your desktop – no payment required. 2. Start processing your audio and ultrasonic recordings automatically. 3. Review the status and see the results on your management dashboard.
Scalable pricing and usage
The Pipeline is free, subject to fair usage and data sharing conditions. Pay-as-you-go and bespoke paid Pipeline Projects are available for commercial and large-scale use, and provide extra data management options.
Get started
It’s quick and easy. Just fill out a short form and download the app. No payment card is required. Start with the free plan and upgrade as you go. We handle all your data in line with our Terms and Conditions and Privacy Policy.
Features and benefits
Detection and identification of wildlife in ultrasonic and audible sound recordings
Ultrasonic recordings

Our superior algorithms can identify bats, small mammals, bush-crickets and moths.
- More about the Pipeline’s ultrasonic classifiers.
Separate identification of bat echolocation, social and feeding buzz calls
- Learn more about this feature on the Ultrasonic classifiers page.
- Or read the new Acoustic Pipeline blog post, Unlocking the secrets of European bats with the BTO Acoustic Pipeline.
Identification of important regional and island variation in European bats
- Pipistrellus pygmaeus (Iberia), Pipistrellus pipistrellus (Malta), Rhinolophus hipposideros minimus (Malta, North Africa), Pipistrellus crypticus - not official name.
Audible sound recordings

We offer bespoke classifiers for several key bird species and groups.
From single-species classifiers for Nightjar and Eurasian Curlew to multi-species classifiers for nocturnal migrants and European breeding birds, learn more about our exciting work to develop these tools.
- More about the Pipeline's audible classifiers.
Secure, reliable, automated processing
The Pipeline uses cutting-edge machine learning techniques and cloud computing to automate the processing of audible and ultrasonic audio data, providing a platform that can scale to accommodate acoustic monitoring projects of any size.
More about how the Acoustic Pipeline works.
Advanced identfication algorithms
The species identification algorithms in the Pipeline deliver unique and unrivalled performance for birds, bats and other wildlife. Developed by species experts Dr Stuart Newson and Dr Simon Gillings in collaboration with other experts throughout Europe, distilling years of audio recording and first-hand experience.
Trusted by thousands of people
Since its launch in 2021, the BTO Acoustic Pipeline has processed over 500 terabytes of audio recordings from more than 2000 users in 25 countries in Europe and beyond. It has supported county councils, National Parks, rewilding projects and NGOs to audit their biodiversity.
Find our more, read our Case Studies.
For commercial use, academic research and citizen science
The Pipeline has been used for a wide range of commercial projects, citizen science projects and bioacoustics research. It is trusted by over 100 ecological consultancies including some of the biggest players in the UK, enhancing the quality of data used to inform planning decisions.
Find our more about commercial and large-scale use.
Free support from our in-house experts, from start to finish

Our Acoustic Pipeline Support Hub is available for free before you even begin recording. It provides a range of guides and advice to ensure a smooth and easy end-to-end workflow for acoustic monitoring. Support from our in-house experts Dr Simon Gillings and Dr Stuart Newson is also available from acoustic.pipeline@bto.org
Questions?
We’d love to hear from you – just get in touch
Contact us by email at acoustic.pipelinebto.org
You can also learn more about and contact the scientists behind the Pipeline:
Latest Pipeline projects
Monitoring habitat restoration work
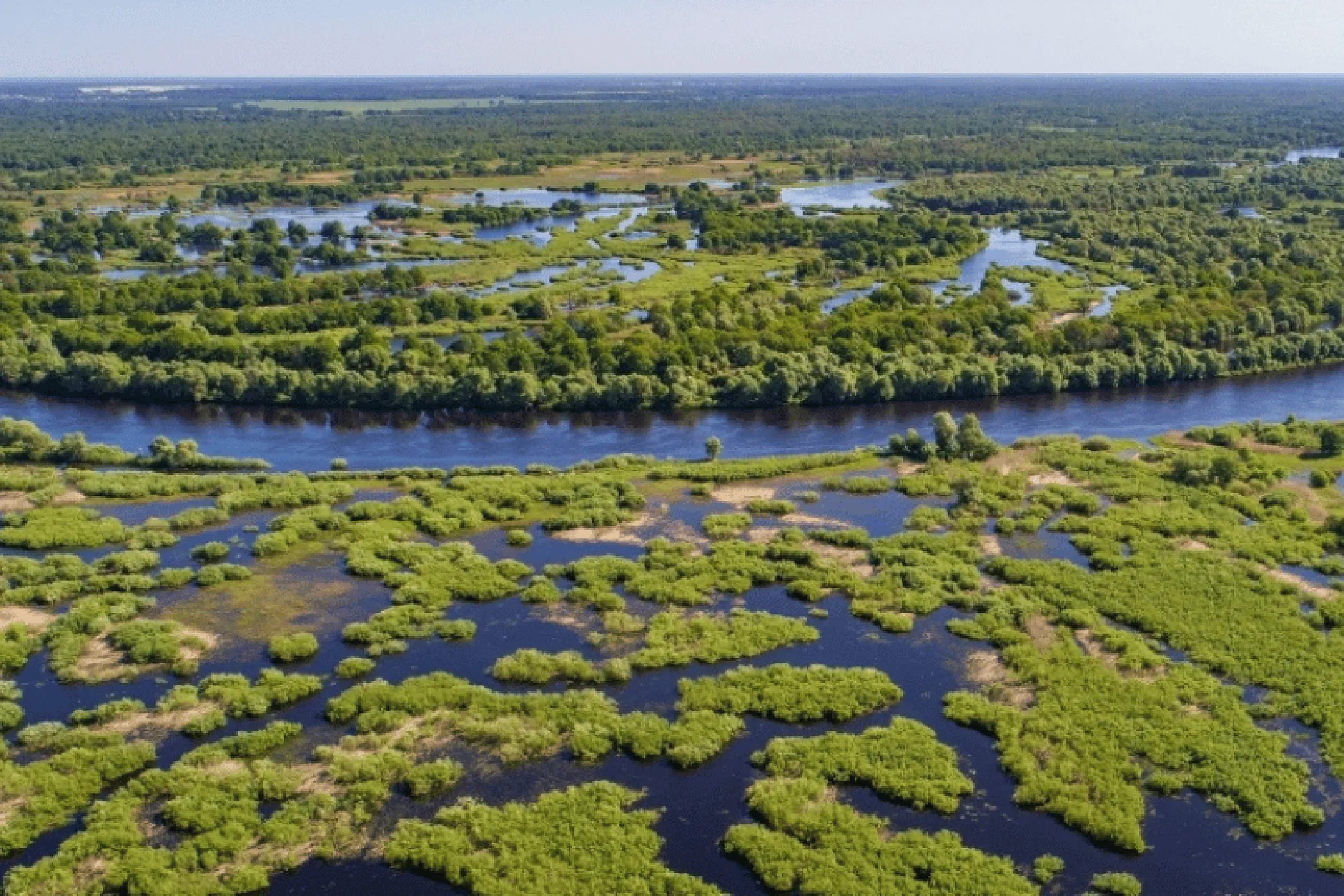
As part of the Endangered Landscapes Programme, the Pipeline processed recordings from over 400 locations across Polesia.

Recording nocturnal bird migration
We trained a neural network to detect and identify flight calls of nocturnally migrating birds for a peer-reviewed publication.

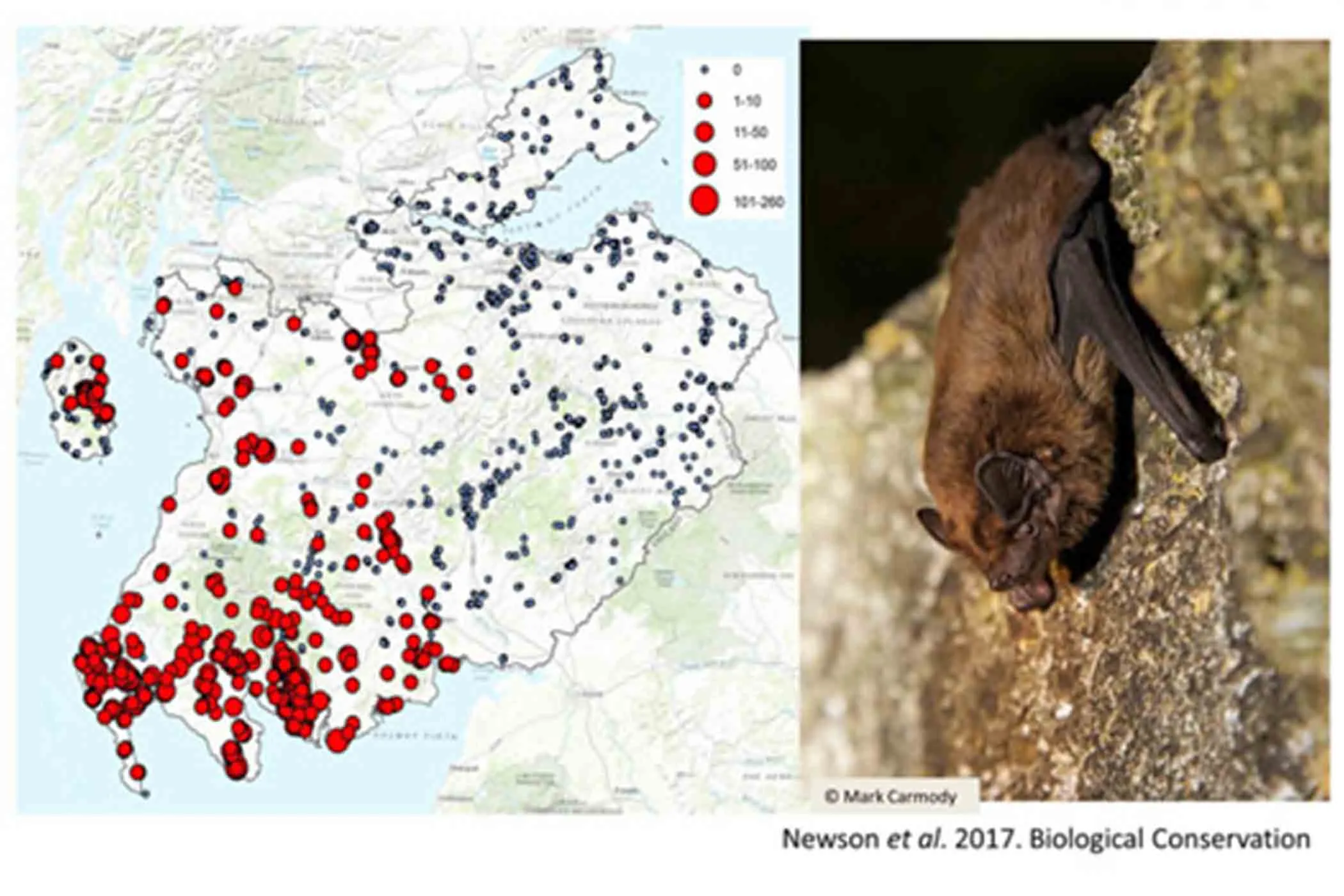
Quantifying the impact of wind farms on bats
Acoustic Pipeline classifiers processed recordings from over 20,000 km2 of southern Scotland to identify the top areas for bats.
Choose from free or paid plans, and bespoke projects
We have three plans available: free, pay as you go and bespoke.
The Pipeline is provided for free, subject to fair usage and data sharing conditions. We also offer a range of scalable, pay-as-you-go plans for greater processing capacity and for when data sharing is not possible.
For commercial or large-scale use, we offer a bespoke service called Acoustic Pipeline Projects. This service provides extra tools for data management as well as the Pipeline’s species identification capabilities.
Get started with the Pipeline
It’s quick and easy to register for the Pipeline. No payment card is required.
Terms, Conditions and Privacy
We take your privacy seriously, and handle all your data in line with the following policies:
Get in touch
Interested in a Pipeline Project?
We’d love to hear from you.
Email us at acoustic.pipeline@bto.org
Acknowledgement
The development of the BTO Acoustic Pipeline infrastructure and its classifiers has been supported by various grants, research awards, charitable donations and contracts. In particular, we would like to thank the Esmée Fairbairn Foundation, Ken and Linda Smith, Natural England, and the Endangered Landscapes Programme, which is managed by the Cambridge Conservation Initiative in partnership with Arcadia.
